Methylation and UW-RUA
Mussel Methylation
Planning
Lab Notebook
Plan of the Week: April 6 - April 12, 2026
High- level outline for the week. Adjusted daily to reflect progress of the day before
- This week’s plan is to continue to move forward with the methylation analysis, refresh mutual goals, expectations and priorities with Steven, and set goals for April.
Monday - Planning the week & Setting April Goals
Tuesday - UW-RUA, No Science
Wednesday - Friday: Will Outline post Steven 1v1
Saturday - No Science
Sunday - Reading
Plan of the Day
Granular level task list to accomplish the high- level goal outlined above
- Continue to move forward with the methylation analysis.
- Knockout UW-RUA tasks, including traveler management
Projects Touched Today
- DNA Methylation
Progress Notes
- Today’s first priority was checking in on the methylation extractions; they were almost finished last night.
- All sequences were completed by 0500. All reports from Bismark and MultiQC were completed by 0600.
- My rsync problem was that I was trying to run it from my code directory… I knew I just needed to step away before trying again. I was able to get that running before Apple decided my computer update was more important - so I restarted it after being disconnected, but it is decidedly slower than molasses in January so we wait.
- Next up is to commit the smaller files to GH and start to look at the reports and ask questions.
- Next, I updated my notebook posts from the weekend. I started them each day, but didn’t finish them.
- Once that was done, I had to shift to some UW-RUA traveler management tasks that are time sensitive. My next step is April goal planning and building my 1v1 agenda for Wednesday.
- I took a look at the methylation extraction reports from Bismark and MultiQC. A couple things I jotted down while reviewing:
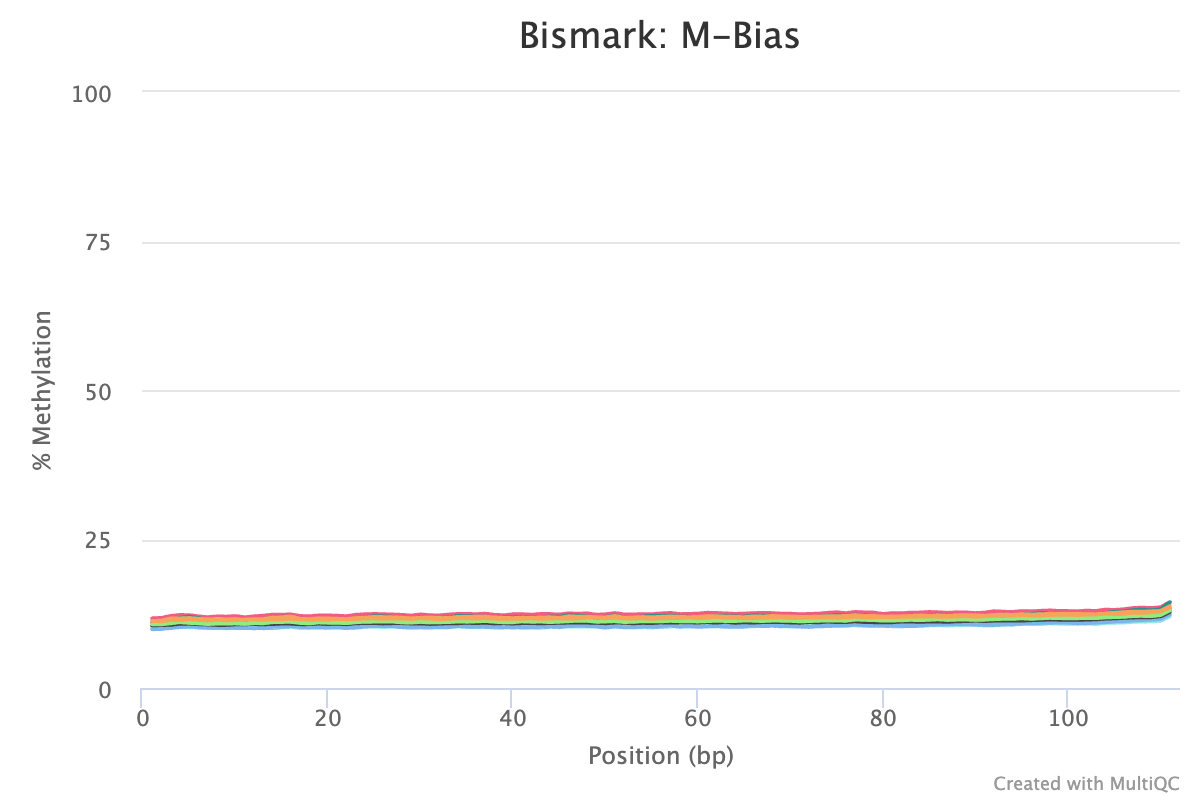
- M-Bias reports/ plots. ‘M’ethylation bias across read positions. M-bias looks at the percentage of cytosines are ’called’ methylated across every single base in the sequence. Additionally, the lack of any ‘jumps’ or ‘spikes’ in the lines indicate nothing external (non-biological) has created noise in the sequences.
- On average, all sequences are around 11% methylation at CpGs (see sample 272M plot from Bismark below). This means things like the trimming parameters and bisulfite conversion are consistent and not driving the methylation percentage up or down from a process POV.
- M-Bias reports/ plots. ‘M’ethylation bias across read positions. M-bias looks at the percentage of cytosines are ’called’ methylated across every single base in the sequence. Additionally, the lack of any ‘jumps’ or ‘spikes’ in the lines indicate nothing external (non-biological) has created noise in the sequences.
- Since I like a good list over a plot, I took the multiQC data table, added pertinent sample information and put it below. It helped me see that methylation, on average was the same across all samples - that makes the upcoming work of figuring out where that methylation is occurring very interesting.
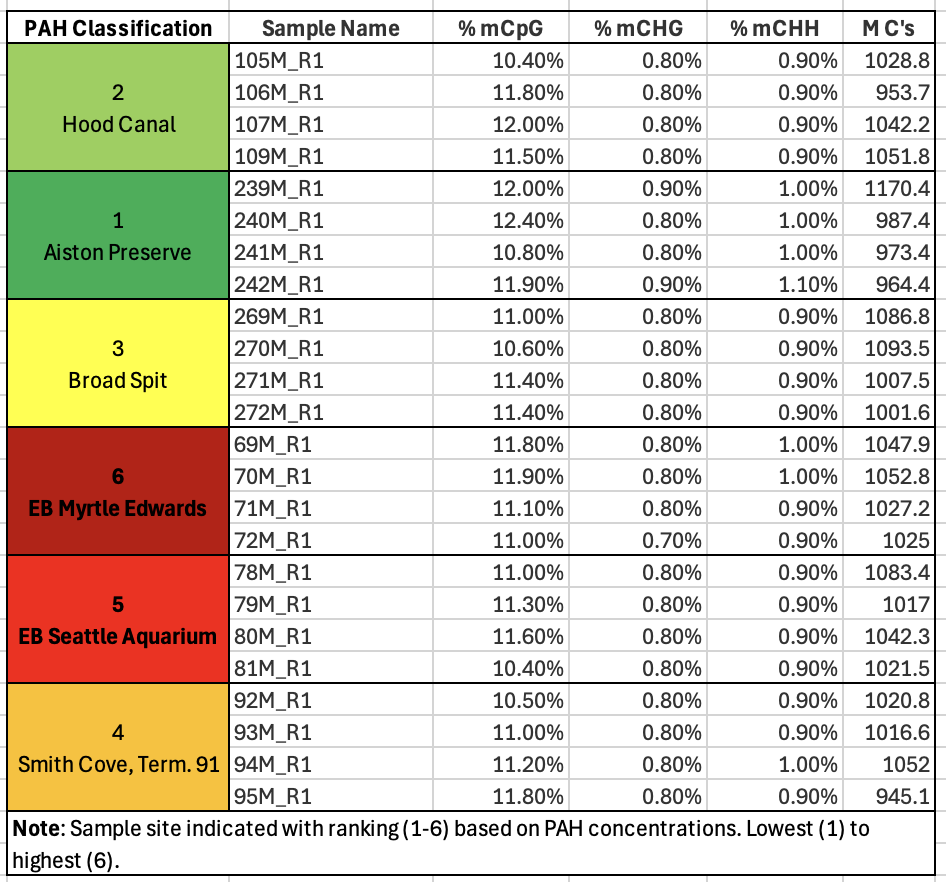
Additionally, the low percentages of methylation at CHG or CHH (any cytosine that is not followed by a G(uanine) directly are low. This is important because what we know about animal methylation and it’s potential functional relevance tells us CpGs are where it’s at, so high numbers could indicate a bisulfite conversion issue in the data.
- Fun fact, methylation in plants typically occurs at CG, CHG or CHH sites. Check out this Muyle et al. paper from 2022 that explains it way better than me!
After looking over my reports, I returned to UW-RUA to prep for tomorrow.
Outcomes: Products & Word Count
- Something: 0 words
Today’s total: 0 words
Monthly total to date: 921 words
Annual total to date: 33,593 words
Annual target total to date: 47,500 words
Next Up: Tomorrow’s Plan
- Wrapping up my agenda for my 1v1 with Steven, setting up some April goals, and of course, UW-RUA.